Analysing and plotting magnetic resonance data¶
This notebook contains a quick example of plotting data stored in magres files, without performing any referencing.
[1]:
from matador.scrapers import magres2dict
from matador.plotting import plot_magres
magres, failures = magres2dict("*.magres", as_model=True)
Successfully scraped 2 out of 2 files.
[2]:
for doc in magres:
print(doc)
doc.print_sites()
Li3P: LiP_CASTEP18.magres
=========================
20 atoms. P1
(a, b, c) = 4.1333 Å, 6.0638 Å, 12.5450 Å
(α, β, γ) = 88.4518° 80.5130° 90.0085°
---
0 Li 0.0916 0.3584 0.9424
chemical_shielding_iso = 83.7003
magnetic_shielding_tensor =
[[ 8.16e+01 6.11e-03 -4.30e-02]
[ 1.27e-02 7.99e+01 1.36e+00]
[-5.07e-02 2.13e+00 8.96e+01]]
chemical_shift_aniso = 9.3772
chemical_shift_asymmetry = 0.3284
---
1 Li 0.1983 0.7851 0.7195
chemical_shielding_iso = 84.3395
magnetic_shielding_tensor =
[[8.40e+01 9.13e-02 4.66e-02]
[1.30e-01 8.70e+01 9.94e-01]
[9.87e-02 1.57e+00 8.21e+01]]
chemical_shift_aniso = 4.4293
chemical_shift_asymmetry = 0.7621
---
2 Li 0.2807 0.9980 0.5589
chemical_shielding_iso = 83.3730
magnetic_shielding_tensor =
[[ 8.12e+01 -1.51e-02 -4.42e-02]
[ 1.19e-02 8.44e+01 -4.39e+00]
[ 2.60e-03 -4.26e+00 8.45e+01]]
chemical_shift_aniso = 8.1433
chemical_shift_asymmetry = 0.1863
---
3 Li 0.3473 0.7369 0.4079
chemical_shielding_iso = 86.6380
magnetic_shielding_tensor =
[[ 8.52e+01 4.23e-02 -2.41e-02]
[ 4.89e-02 8.82e+01 1.01e+00]
[-2.43e-03 7.34e-01 8.65e+01]]
chemical_shift_aniso = 2.9372
chemical_shift_asymmetry = 0.4602
---
4 Li 0.4349 0.4794 0.2566
chemical_shielding_iso = 83.3435
magnetic_shielding_tensor =
[[81.2 0.13 -0.17]
[ 0.13 84.39 -4.39]
[-0.09 -4.27 84.44]]
chemical_shift_aniso = 8.1073
chemical_shift_asymmetry = 0.2058
---
5 Li 0.4413 0.0210 0.2380
chemical_shielding_iso = 85.0509
magnetic_shielding_tensor =
[[ 8.30e+01 8.90e-02 -4.87e-02]
[ 1.22e-01 8.63e+01 1.20e+00]
[ 4.24e-04 1.24e+00 8.59e+01]]
chemical_shift_aniso = 3.4360
chemical_shift_asymmetry = 0.8373
---
6 Li 0.5173 0.6897 0.0948
chemical_shielding_iso = 84.4205
magnetic_shielding_tensor =
[[8.39e+01 4.68e-02 7.09e-02]
[1.32e-01 8.73e+01 1.02e+00]
[8.51e-02 1.60e+00 8.21e+01]]
chemical_shift_aniso = 4.7370
chemical_shift_asymmetry = 0.6511
---
7 Li 0.6253 0.1122 0.8730
chemical_shielding_iso = 83.7612
magnetic_shielding_tensor =
[[ 8.17e+01 1.11e-02 -1.06e-01]
[ 4.81e-03 8.00e+01 1.34e+00]
[-1.34e-01 2.11e+00 8.95e+01]]
chemical_shift_aniso = 9.1321
chemical_shift_asymmetry = 0.3245
---
8 Li 0.6906 0.4626 0.7373
chemical_shielding_iso = 82.8192
magnetic_shielding_tensor =
[[ 7.98e+01 1.35e-01 -7.89e-02]
[ 1.16e-01 8.67e+01 -5.36e+00]
[-8.88e-02 -3.92e+00 8.19e+01]]
chemical_shift_aniso = 10.0677
chemical_shift_asymmetry = 0.1121
---
9 Li 0.7305 0.0937 0.6585
chemical_shielding_iso = 83.6264
magnetic_shielding_tensor =
[[ 8.39e+01 -2.15e-02 -2.34e-02]
[-1.26e-02 8.57e+01 5.72e+00]
[-5.97e-02 5.90e+00 8.13e+01]]
chemical_shift_aniso = -9.5122
chemical_shift_asymmetry = 0.9216
---
10 Li 0.8485 0.4570 0.4171
chemical_shielding_iso = 84.9298
magnetic_shielding_tensor =
[[ 9.07e+01 1.70e-01 -6.90e-02]
[ 1.65e-01 8.44e+01 1.00e+00]
[ 8.56e-03 4.94e-01 7.97e+01]]
chemical_shift_aniso = 8.6833
chemical_shift_asymmetry = 0.8534
---
11 Li 0.8585 0.0210 0.3974
chemical_shielding_iso = 84.9095
magnetic_shielding_tensor =
[[ 9.08e+01 -8.07e-02 -9.23e-02]
[-5.25e-02 8.44e+01 1.10e+00]
[-2.69e-02 5.81e-01 7.96e+01]]
chemical_shift_aniso = 8.7771
chemical_shift_asymmetry = 0.8596
---
12 Li 0.9853 0.3823 0.1564
chemical_shielding_iso = 83.6033
magnetic_shielding_tensor =
[[ 8.39e+01 -8.91e-03 -3.55e-02]
[-2.42e-03 8.57e+01 5.72e+00]
[-8.39e-02 5.95e+00 8.12e+01]]
chemical_shift_aniso = -9.5887
chemical_shift_asymmetry = 0.9116
---
13 Li 0.0239 0.0140 0.0804
chemical_shielding_iso = 82.6785
magnetic_shielding_tensor =
[[ 7.96e+01 1.38e-01 -8.10e-02]
[ 1.13e-01 8.66e+01 -5.51e+00]
[-1.02e-01 -4.10e+00 8.18e+01]]
chemical_shift_aniso = 10.3680
chemical_shift_asymmetry = 0.1147
---
14 Li 0.2721 0.4547 0.5780
chemical_shielding_iso = 85.0693
magnetic_shielding_tensor =
[[ 8.29e+01 1.08e-01 3.27e-03]
[ 1.34e-01 8.64e+01 1.12e+00]
[-5.75e-03 1.17e+00 8.59e+01]]
chemical_shift_aniso = 3.3677
chemical_shift_asymmetry = 0.9200
---
15 P 0.1818 0.2116 0.7570
chemical_shielding_iso = 350.0256
magnetic_shielding_tensor =
[[ 8.80e+01 5.62e-01 -3.10e-01]
[-8.16e-01 5.39e+02 9.52e+00]
[-9.03e-01 -6.02e+01 4.23e+02]]
chemical_shift_aniso = -392.9941
chemical_shift_asymmetry = 0.4806
---
16 P 0.3536 0.2387 0.4077
chemical_shielding_iso = 500.3197
magnetic_shielding_tensor =
[[497.42 10.84 12.11]
[ 0.66 418.21 -66.52]
[ 3.68 -66.28 585.33]]
chemical_shift_aniso = 162.6815
chemical_shift_asymmetry = 0.9530
---
17 P 0.5341 0.2630 0.0584
chemical_shielding_iso = 353.3182
magnetic_shielding_tensor =
[[ 9.25e+01 5.45e-01 1.54e+00]
[-1.74e-01 5.40e+02 1.12e+01]
[-9.61e-01 -5.78e+01 4.27e+02]]
chemical_shift_aniso = -391.2453
chemical_shift_asymmetry = 0.4668
---
18 P 0.7688 0.7316 0.5822
chemical_shielding_iso = 530.9353
magnetic_shielding_tensor =
[[ 4.13e+02 8.13e+00 4.95e+00]
[ 6.32e-01 5.05e+02 -9.77e+00]
[ 1.06e+00 -4.89e+01 6.75e+02]]
chemical_shift_aniso = 223.9065
chemical_shift_asymmetry = 0.5876
---
19 P 0.9456 0.7440 0.2335
chemical_shielding_iso = 531.2349
magnetic_shielding_tensor =
[[411.31 8.65 7.53]
[ 0.72 505.85 -7.57]
[ 1.75 -45.03 676.55]]
chemical_shift_aniso = 223.9919
chemical_shift_asymmetry = 0.6112
---
Na3P: NaP_QE6.magres
====================
4 atoms. Fm-3m
(a, b, c) = 5.0075 Å, 5.0075 Å, 5.0075 Å
(α, β, γ) = 120.0020° 120.0020° 89.9965°
---
0 Na 0.0000 0.0000 0.0000
chemical_shielding_iso = 518.1528
magnetic_shielding_tensor =
[[518.53 -0. 0. ]
[ -0. 518.53 0. ]
[ -0. 0. 517.4 ]]
chemical_shift_aniso = -1.1365
chemical_shift_asymmetry = -0.0000
---
1 Na 0.2500 0.7500 0.5000
chemical_shielding_iso = 467.6065
magnetic_shielding_tensor =
[[468.02 -0. 0. ]
[ -0. 468.02 0. ]
[ 0. 0. 466.79]]
chemical_shift_aniso = -1.2261
chemical_shift_asymmetry = -0.0000
---
2 Na 0.7500 0.2500 0.5000
chemical_shielding_iso = 467.6065
magnetic_shielding_tensor =
[[468.02 0. -0. ]
[ 0. 468.02 0. ]
[ -0. 0. 466.79]]
chemical_shift_aniso = -1.2261
chemical_shift_asymmetry = -0.0000
---
3 P 0.5000 0.5000 0.0000
chemical_shielding_iso = 275.3424
magnetic_shielding_tensor =
[[273.7 -0. -0. ]
[ -0. 273.7 0. ]
[ -0. 0. 278.62]]
chemical_shift_aniso = 4.9146
chemical_shift_asymmetry = 0.0000
---
Species as separate figures¶
[3]:
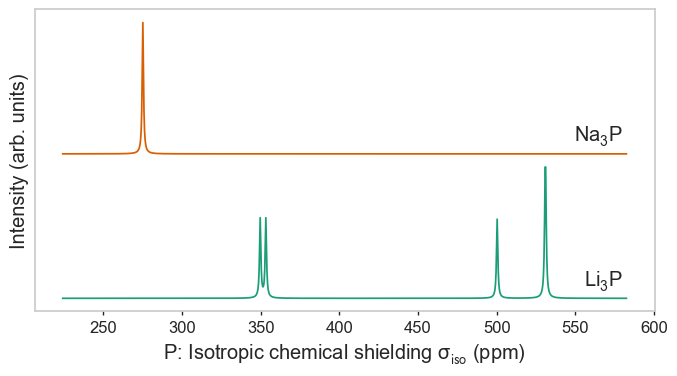
plot_magres(magres, "P")
plot_magres(magres, "Na")
plot_magres(magres, "Li", broadening_width=0.01);
No sites of Na found in LiP_CASTEP18.magres, signal will be empty.
No sites of Li found in NaP_QE6.magres, signal will be empty.


Species as subplots with custom colours and labels¶
[4]:
import matplotlib.pyplot as plt
fig, axes = plt.subplots(2, 1, figsize=(8, 6))
line_kwargs = [{"c": "red", "ls": "--"}, {"c": "black", "ls": "-."}]
labels = ["B", "A"]
plot_magres(
magres, "Na",
ax=axes[0], line_kwargs=line_kwargs, signal_labels=labels
)
plot_magres(
magres, "Li", broadening_width=0.01,
ax=axes[1], line_kwargs=line_kwargs, signal_labels=labels
)
plt.tight_layout()
No sites of Na found in LiP_CASTEP18.magres, signal will be empty.
No sites of Li found in NaP_QE6.magres, signal will be empty.

Other magres quantities as subplots¶
[5]:
import matplotlib.pyplot as plt
fig, axes = plt.subplots(2, 1, figsize=(8, 6))
plot_magres(
magres, "P", magres_key="chemical_shift_aniso",
ax=axes[0]
)
plot_magres(
magres, "P", magres_key="chemical_shift_asymmetry",
ax=axes[1],
broadening_width=0,
)
plt.tight_layout()
